Repub Eur Nl
18/8/14 Epidemiological studies have shown that consumption of a highfat diet (HFD) increases the risk of developing breast cancer (BC) Studies in rodents have shown HFD causes changes in the genetic programming of the maturing mammary gland (MG) increasing the susceptibility of developing the disease Less is known about how HFD induced genes impact BCCHARAFE_BREAST_CANCER_BASAL_VS_MESENCHYMAL_UP # features = 14 chi−square p = 08 CHARAFE_BREAST_CANCER_BASAL_VS_MESENCHYMAL_UP # features = 14 , max = 2
Charafe_breast_cancer_luminal_vs_ mesenchymal up
Charafe_breast_cancer_luminal_vs_ mesenchymal up-In breast cancer the presence of cells undergoing the epithelialtomesenchymal transition is indicative of metastasis progression Since metabolic features of breast tumour cells are critical in cancer progression and drug resistance, we hypothesized that the lipid content of malignant cells might be a useful indirect measure of cancer progressionFigures 2, 5, and 6 Contribute to jrboyd/paper_v5 development by creating an account on GitHub
Repub Eur Nl
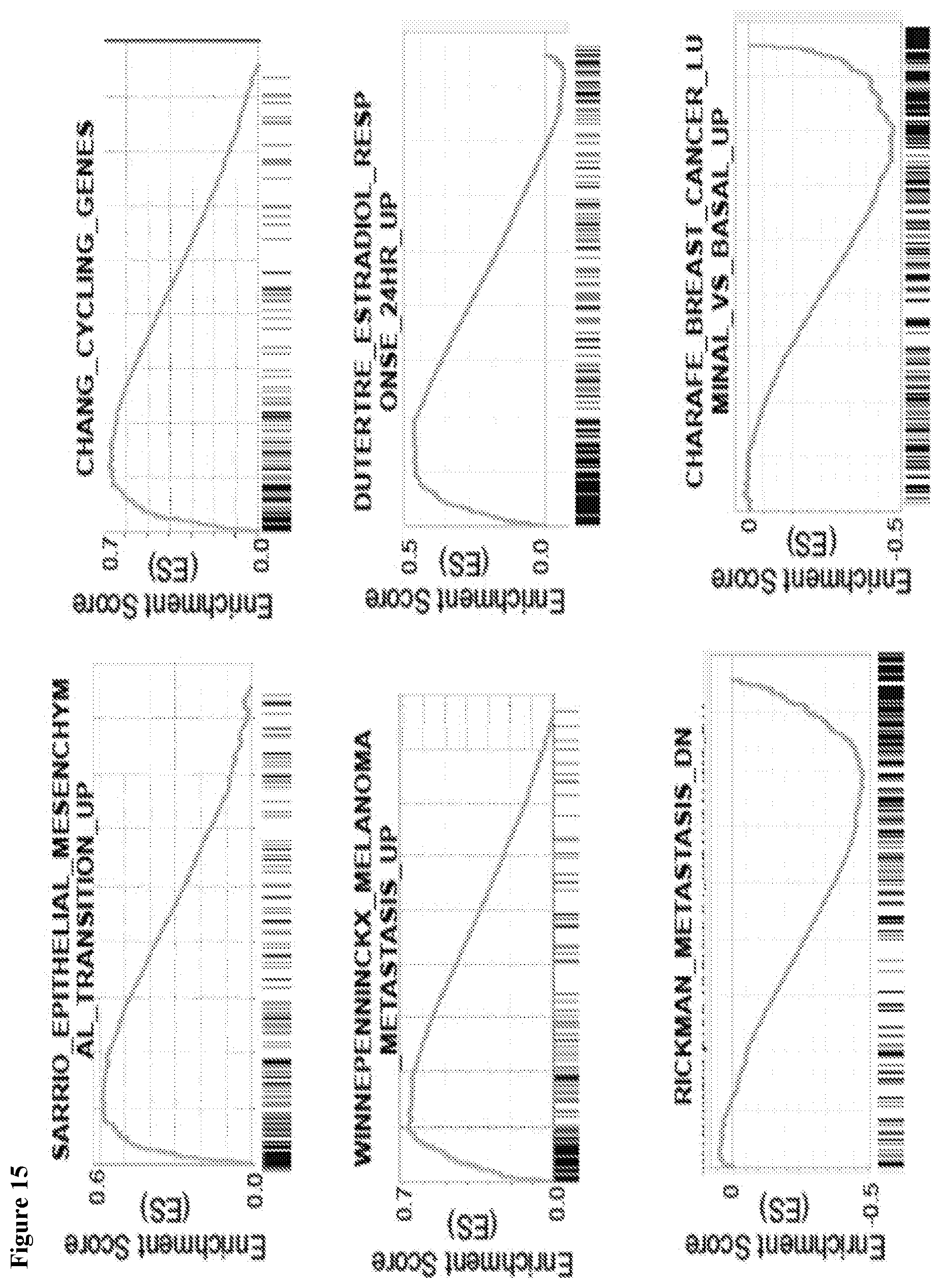
9/8/19 Breast cancer is the most common cancer among women, and although targeted therapies have helped to significantly reduced breast cancer mortality rate in the past decade, further improvement will require a comprehensive understanding of the molecular mechanisms of the disease (1, 2)Recently, deep mass spectrometry based proteomic characterization of1/9/14 As expected, MIII cells showed a more mesenchymal phenotype, with significant gene sets being those genes described by Charafe et al as downregulated in basal and luminal breast cancer subtypes According to these transcriptomic public data, we inferred that fatty acid and glucose metabolic functions differentiate confluent and sparse MCF10A cells, then supporting1/3/13 Furthermore, a geneset associated with the luminal phenotype (Charafe Breast Cancer Luminal vs Mesenchymal Up) was also enriched in ESUM149 clones Conversely, genesets associated with the loss of ECadherin or mesenchymal phenotype were enriched in MSUM149 clones (Table (Table1, 1, Fig Fig3c) 3c)
Charafe breast cancer basal vs mesenchymal up entpd2, epb41l4a, fgfbp1, maf, ppl, rnf144b, sh2d3a, slc11a2, sytl1, tc2n module 4 btg2, cxcl1, fgfbp1, gas1, il6 thiolester hydrolase activity acot11, acot8, otub2, usp53 gcm map4k4 charafe breast cancer luminal vs mesenchymal upCharafe_breast_cancer_luminal_vs_mesenchy mal_dn 0 0 459 0481 gao_large_intestine_adult_cj_immune_cells 0 0 502 0514 onder_cdh1_targets_2_up 0 0 250 0556 aizarani_liver_c21_stellate_cells_1 0 0 192 05 6411Other studies in breast cancer have shown that mir106b/25 cluster activates TGFβ signaling and epithelialmesenchymal transition (EMT) and miR221/222 cluster is a key regulator of luminal breast cancer tumor progression
Charafe_breast_cancer_luminal_vs_ mesenchymal upのギャラリー
各画像をクリックすると、ダウンロードまたは拡大表示できます
 |  |  |
 |  | |
 | ||
「Charafe_breast_cancer_luminal_vs_ mesenchymal up」の画像ギャラリー、詳細は各画像をクリックしてください。
 |  | |
 |  | |
 |  | |
「Charafe_breast_cancer_luminal_vs_ mesenchymal up」の画像ギャラリー、詳細は各画像をクリックしてください。
 | ||
 |  |  |
 | ||
「Charafe_breast_cancer_luminal_vs_ mesenchymal up」の画像ギャラリー、詳細は各画像をクリックしてください。
 | ||
 |  | |
 |  | |
「Charafe_breast_cancer_luminal_vs_ mesenchymal up」の画像ギャラリー、詳細は各画像をクリックしてください。
 |  |  |
 |  |  |
 | ||
「Charafe_breast_cancer_luminal_vs_ mesenchymal up」の画像ギャラリー、詳細は各画像をクリックしてください。
 |  |  |
 | ||
「Charafe_breast_cancer_luminal_vs_ mesenchymal up」の画像ギャラリー、詳細は各画像をクリックしてください。
 |  |  |
 | ||
 |  | |
「Charafe_breast_cancer_luminal_vs_ mesenchymal up」の画像ギャラリー、詳細は各画像をクリックしてください。
 |  | |
 | ||
「Charafe_breast_cancer_luminal_vs_ mesenchymal up」の画像ギャラリー、詳細は各画像をクリックしてください。
 |  | |
 |  | |
 |  | |
「Charafe_breast_cancer_luminal_vs_ mesenchymal up」の画像ギャラリー、詳細は各画像をクリックしてください。
 |  | |
 |  | |
 | ||
「Charafe_breast_cancer_luminal_vs_ mesenchymal up」の画像ギャラリー、詳細は各画像をクリックしてください。
 |  |  |
 |  | |
 |  | |
「Charafe_breast_cancer_luminal_vs_ mesenchymal up」の画像ギャラリー、詳細は各画像をクリックしてください。
 |
Motivation Although many gene set analysis methods have been proposed to explore associations between a phenotype and a group of genes sharing common biological functions or involved in the same biological process, the underlying biological mechanisms of identified gene sets are typically unexplained Results We propose a method called Differential Regulationbased enrichmentFigures 2, 5, and 6 Contribute to jrboyd/paper_v5 development by creating an account on GitHub
Incoming Term: charafe_breast_cancer_luminal_vs_ mesenchymal up,




0 件のコメント:
コメントを投稿